by Marjory Moreau, Ph.D.
I have worked in PBPK modeling for many years and until a few months ago, I never had to deal with the brain compartment. Well, I had a brain compartment in several of my models, but I never really thought about the blood brain barrier and all the transporters involved in protecting or bringing compounds into the brain until recently. I remember my mentor telling me about the parsimony principal and keeping my model as simple as possible. Well, sometimes, simple does not work, and we need to think more about what is really happening in an organ or tissue. When it comes to the brain, that becomes even more interesting!
If you read my precedent blogs, you know that PBPK modeling is an efficient and reliable tool to predict the distribution and fate of drugs within an organism. PBPK models include biologically realistic descriptions of tissues and processes involved in the absorption, distribution, metabolism, and excretion (ADME) of a compound. Used in combination with state-of-the-art quantitative structure-activity relationship (QSAR), read-across, and in vitro testing technologies, PBPK models can predict and assess drug efficacy and safety or chemical toxicity by providing estimation of local concentrations at the site of action. PBPK models are complex and depend on many parameters. The PBPK model parameters are divided into three categories, the physiological parameters specific to each species (e.g., age, weight, tissue volumes, blood flows), the drug specific physicochemical parameters (e.g., permeability through membranes, partitioning to tissues, binding to plasma proteins, or affinities towards certain enzymes and transporter proteins, hepatic or renal clearances) and the scenario of exposure (e.g., dose, route and frequency of administration).
The brain is a complex compartment with the blood brain barrier (BBB) serving as a gatekeeper for delivery to the central nervous system (CNS). For compounds with expected neurological activity, it is therefore important to consider the relationship between exposure and brain concentrations. PBPK modeling of the CNS provides the opportunity to predict relevant compound concentrations at the target site. The brain compartment in a PBPK model can be relatively simple or very complex by accounting for known anatomy and physiology of the brain, including the presence of distinct blood-brain barrier and active transporters (Figure 1). The CNS consists of the brain tissue, the surrounding extracellular fluid (ECF) and the cerebrospinal fluid (CSF), as well as the barriers separating these structures from the systemic circulation, the BBB and the blood cerebrospinal fluid barrier (BCSFB).
Qbrain represents the blood flow to the brain, PSb, PSC and PSE represents the passive permeability-surface area products for the three barriers, BBB, BCSFB and brain-CSF barrier, respectively. CLbin and CLin represent the overall clearance of influx transporters expressed at the BBB and BCSFB. CLmet is the metabolic clearance due to brain enzymes. Qsin and Qsout represent the CSF shuttle flow rate from the cranial to the spinal part and vice versa. Qcsink and Qssink represent the CSF flow out of the cranial and spinal CSF compartments, respectively.
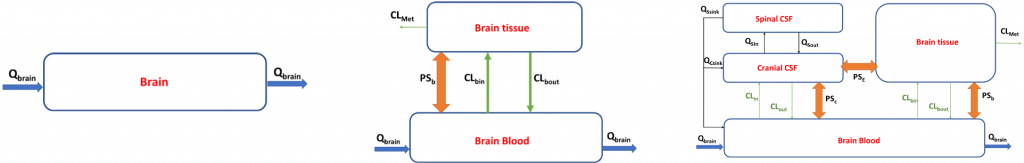
Figure 1. Brain representations in a PBPK Model (Adapted from Ball et al. 2013)
In PBPK modeling, tissues can be represented as perfusion or permeability limited compartment. A perfusion limited tissue means that the rate of drug entry into the compartment (tissue) is determined by the organ’s blood flow rate. The compound distributes freely across the membranes without diffusion barriers or significant transporter contributions. For a permeability limited compartment, the limited factor is not the blood flow rate but the membrane permeability and/or transporters kinetic parameters. The brain is considered (most of the time) as permeability limited due to the BBB and its tight junctions and transporters that create a barrier and limit the rate of drug or chemical exchange between blood and brain tissue. If regional distribution of drug or chemical within the brain is expected, a more complex representation of the brain will be required with incorporation of the cerebrospinal fluid compartment and barrier.
Most of these parameters for the brain compartment, as well as clearance data or unbound fraction can be derived from in silico predictions, in vitro experiments with in vitro to in vivo extrapolation (IVIVE) or by fitting the model to in vivo data. Ultimately, the success of the modeling simulations is limited by the quantity and quality of the data available to parametrize and validate the model.
To summarize, with the BBB and the involvement of different type of transporters, PBPK models of the brain are complex. Because almost no human in vivo data are available for the brain tissue and because of species differences that make the scaling to human difficult, the success of PBPK model for prediction of brain pharmacokinetic depends on in vitro systems to estimate the BBB permeability. The brain is an amazing organ and there are so many things to say on this subject that I could continue writing for days! If you are interested in more detail, take a look at these publications and don’t hesitate to contact me: mmoreau@scitovation.com
Ball K, Bouzom F, Scherrmann JM, Walther B, Declèves X. Physiologically based pharmacokinetic modelling of drug penetration across the blood-brain barrier–towards a mechanistic IVIVE-based approach. AAPS J. 2013 Oct;15(4):913-32. doi: 10.1208/s12248-013-9496-0. Epub 2013 Jun 20. PMID: 23784110; PMCID: PMC3787211.
Ball K, Bouzom F, Scherrmann JM, Walther B, Declèves X. Development of a physiologically based pharmacokinetic model for the rat central nervous system and determination of an in vitro-in vivo scaling methodology for the blood-brain barrier permeability of two transporter substrates, morphine and oxycodone. J Pharm Sci. 2012 Nov;101(11):4277-92. doi: 10.1002/jps.23266. Epub 2012 Aug 1. PMID: 22864977.
Gaohua L, Neuhoff S, Johnson TN, Rostami-Hodjegan A, Jamei M. Development of a permeability-limited model of the human brain and cerebrospinal fluid (CSF) to integrate known physiological and biological knowledge: Estimating time varying CSF drug concentrations and their variability using in vitro data. Drug Metab Pharmacokinet. 2016 Jun;31(3):224-33. doi: 10.1016/j.dmpk.2016.03.005. Epub 2016 Apr 4. PMID: 27236639.
Featured Photo by Natasha Connell on Unsplash
Key Takeaways
- PBPK modeling is a powerful tool for predicting drug distribution in the central nervous sytem (CNS), which is important for understanding drug efficacy and safety.
- PBPK models can be used to simulate drug transport across the blood-brain barrier (BBB) and predict the concentrations of drugs in the brain.
- PBPK models can be used to inform drug development and help design safer and more effective CNS drugs.
Ready to explore how ScitoVation can support your project? Contact us today!
Want to learn more? Take a look at:
- Implementing Mode-Of-Action Investigative Toxicology Into Risk Assessments Through New Alternative Approaches
- Pharmacodynamics And Pharmacokinetics Of Drugs In Early Life Stages
- Discovery Of A New Mechanism For Genetic Change: Exitron Splicing Implications For Carcinogenesis
Importance Of PBPK Modeling In Drug Development
Drug development has benefited from the popularity of physiologically based pharmacokinetic (PBPK) modeling, which offers precise predictions of a drug’s pharmacokinetic characteristics, including absorption, distribution, metabolism, and elimination. This modeling aids in assessing drug safety, efficacy, and optimizing dosing regimens.
- PBPK modeling is particularly vital for central nervous system (CNS) drug development, as the blood-brain barrier (BBB) complicates pharmacokinetic predictions. The BBB serves as a protective barrier, limiting drug access to the CNS. PBPK modeling helps understand CNS drug pharmacokinetics and optimize their dosing regimens.
- Additionally, PBPK modeling evaluates drug safety in the CNS, predicting pharmacokinetics to assess potential drug-drug interactions and off-target effects. This ensures drug safety and efficacy before patient administration.
- Lastly, PBPK modeling optimizes dosing regimens by predicting pharmacokinetics in the CNS, ensuring the drug reaches the target tissue at the desired concentration, reducing side effects, and maximizing efficacy.
Also Read: Physiologically Based Pharmacokinetic Modeling (PBPK)
PBPK Modeling Case Studies:
- Case Study: Pyrethroids
- Case Study: Butylparaben
- Case Study: Use Of PBPK Models For Prioritizing Compounds
Components Of A PBPK Model
Physiologically based pharmacokinetic (PBPK) modeling offers valuable insights into drug pharmacokinetics in the central nervous system (CNS). To accurately model a drug’s pharmacokinetics in the CNS, a PBPK model must encompass:
- CNS Physiology: This includes the CNS anatomy, blood-brain barrier (BBB), cerebrospinal fluid (CSF), brain parenchyma, and physiological processes affecting drug movement.
- Pharmacokinetic Model: A mathematical representation of the drug’s pharmacokinetic parameters, including absorption, distribution, metabolism, and elimination.
- Physiologically Based Model: This represents the drug’s pharmacokinetics in the CNS, considering both pharmacokinetic parameters and physiological processes. It also accounts for pharmacodynamic parameters such as drug affinity and elimination rate in the CNS.
- Simulation and Analysis: This component assesses the model’s predictions, analyzing the effects of various drug doses on CNS pharmacokinetics, and evaluating the drug’s safety and efficacy.
Ready to explore PBPK modeling for your CNS drug development? Contact ScitoVation to discuss your project with our experts.
FAQs
What is PBPK modeling?
PBPK modeling is a mathematical modeling technique used to predict the pharmacokinetics (PK) and pharmacodynamics (PD) of drugs in the body. It combines physiological, anatomical and pharmacokinetic data to create a predictive model of drug absorption, distribution, metabolism and excretion.
What is PK vs. PD modeling?
PK modeling is the study of how drugs are absorbed, distributed, metabolized and eliminated from the body. PD modeling is the study of how drugs interact with the body to produce a therapeutic effect.
Which software is used in PK/PD modeling?
There are a variety of software packages available for PK/PD modeling. Some of the most commonly used packages include WinNonlin, NONMEM, Phoenix NLME and Monolix.
What are the uses of PK/PD modeling?
PK/PD modeling can be used to assess the pharmacokinetic and pharmacodynamic properties of drugs, predict drug concentrations in the body, and assess drug interactions. It can also be used to optimize dosing regimens and identify potential drug-drug interactions.
What is PBBM modeling?
PBBM (Physiologically Based Biomarker Modeling) is a type of PK/PD modeling that uses biomarkers to understand the pharmacokinetics and pharmacodynamics of drugs. Biomarkers can be used to measure drug concentrations in the body, predict drug responses, and assess drug interactions.
What does PBPK stand for?
PBPK stands for Physiologically Based Pharmacokinetic modeling.
What is the PBPK approach?
The PBPK approach is a mathematical modeling technique that uses physiological, anatomical and pharmacokinetic data to predict the pharmacokinetics and pharmacodynamics of drugs in the body.
What are the benefits of PBPK modeling?
PBPK modeling can be used to optimize dosing regimens, predict drug concentrations in the body, assess drug interactions, and identify potential drug-drug interactions. It can also be used to assess the safety and efficacy of drugs and to design clinical trials.
What is POP PK vs. PBPK?
POP PK (Population Pharmacokinetic) modeling is a type of PK modeling that uses population data to predict drug concentrations in the body. PBPK (Physiologically Based Pharmacokinetic) modeling is a type of PK modeling that uses physiological, anatomical and pharmacokinetic data to predict drug concentrations in the body.
What are the parameters of PK/PD?
The parameters of PK/PD modeling include drug concentration, clearance rate, volume of distribution, absorption rate, elimination rate, metabolism rate, and pharmacodynamic parameters such as efficacy and potency.
